- About
- Home
- Introduction
- Statistics
- Programs
- Dignet
- Gene
- GenePair
- Help & Docs
- Documents
- Help
- FAQs
- Links
- Acknowledge
- Disclaimer
- Contact Us
Help and Tutorial
This page provides a tutorial for using the features in the Ignet website:
Top right search bar Query
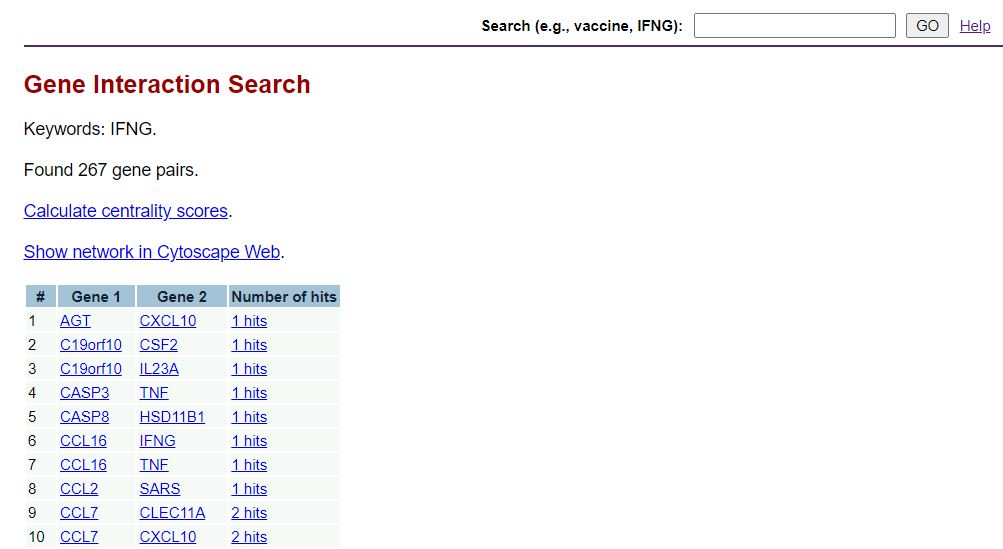
You can type a keywork such as vaccine, IFNG, diabetes. Then click GO. An example is shown here:
Note that the query is indeed the Dignet query. See the screenshot of the results page after the searching for the keyword "IFNG" (e.g., interferon-gamma):
See more information and tutorial about the Dignet query and results description below on this tutorial web page.
Ignet Gene Query
To query a gene, you can choose a gene name, selected whether "vaccine" needs to be mentioned, type a keyword(s) (optional), and then click "Retrieve".
An example is shown below:
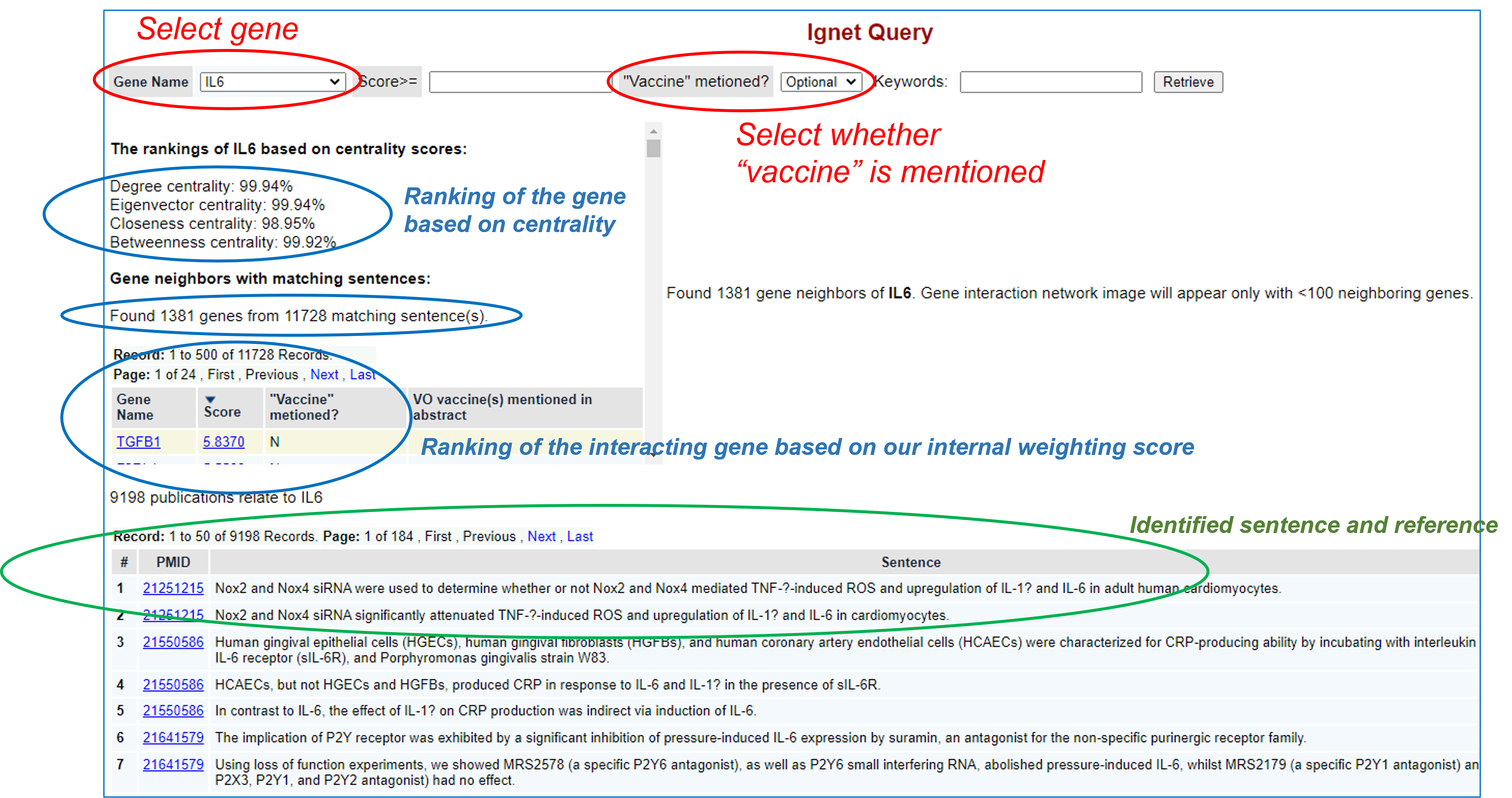
In the example shown in the above screenshot, we selected "IL-6" and left "vaccine" mentioned Optional. See the results below:
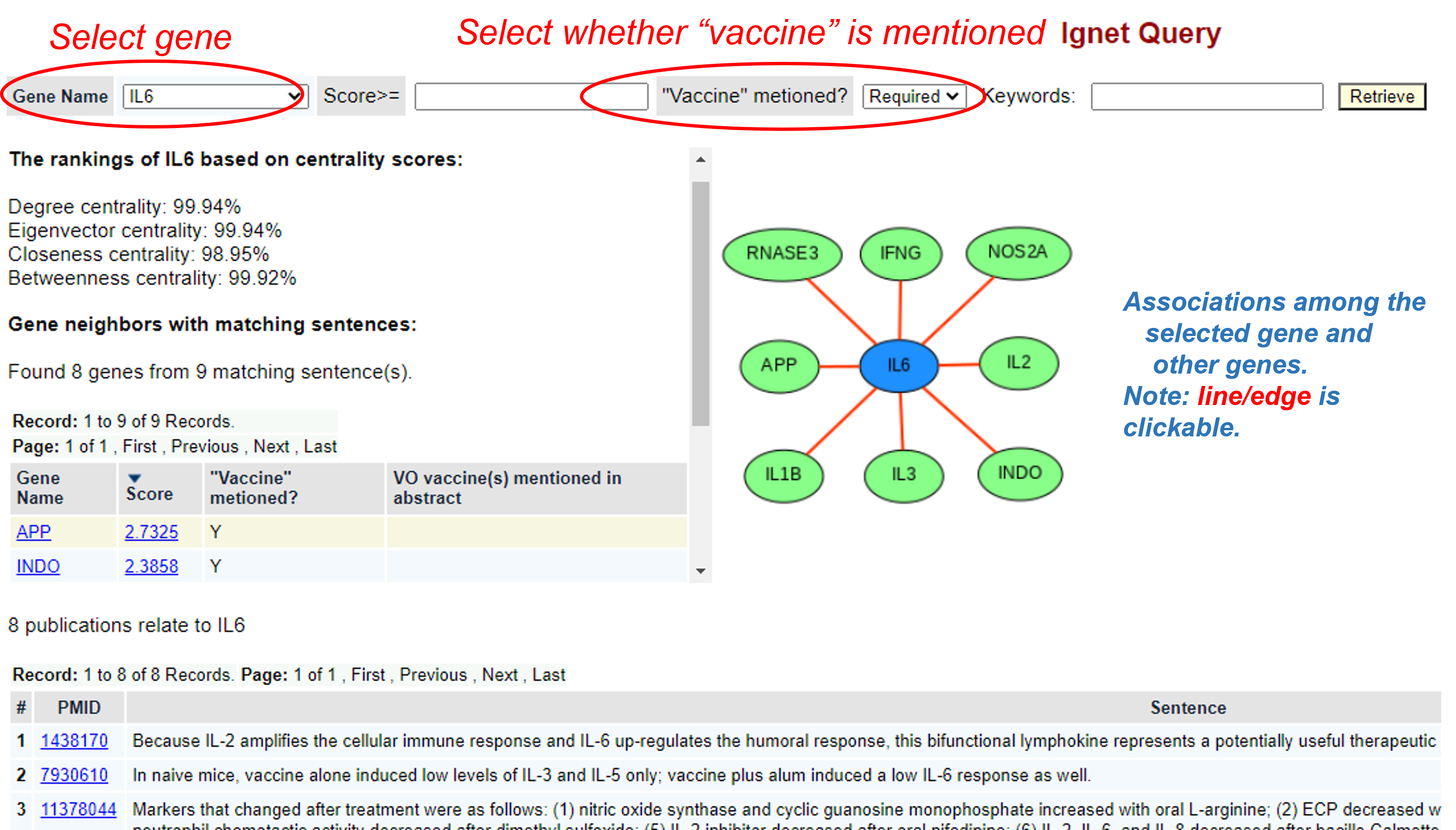

As shown in the above screenshot, the IL6 gene was ranked based on our centraility scores. Four centrality scores, including degree centrality, engenvector centrality, closeness centrality, and betweenness centrality, were calculated. The centrality scores represent the centrality of the IL6 in the identified gene network.
As described in our previous publications (Ozgur et al, 2011, PMC3102897), the meanings of these different types of centralities are:
- Degree centrality: the number of neighbors of a node
- Eigenvector centrality: function of the centralities of a node’s neighbors
- Closeness centrality: inverse sum of the distances from the node to the other nodes in the network
- Betweenness centrality: the proportion of the shortest paths between all the pairs of nodes that pass through the node in interest
In addition, we have also developed an internal weighting score to rank the identified neighboring genes that interact with the target gene. More information will be provided in our journal publication under preparation.
For more examples of Ignet Gene analysis, Choose and Click:
- A1BG, Alpha-1-B Glycoprotein
- IFNG, Interferon-gamma
- IL10, Interleukin-10
- SLC39A1, Solute Carrier Family 39 Member 1
- TLR4, Toll-like recepter 4
- TNF, Tumor necrosis factor
Note: For each of the query, you can add the requirement of 'vaccine' mentioned, and/or you can add some keyword(s) such as 'blood'.
Ignet GenePair Query
Basically, on the GenePair page, you can choose two genes, and they type one keyword(s) such as blood (optional), and then click "Search" to generate the results:
For example, you can choose IL6 and IFNG, and then click "Search":
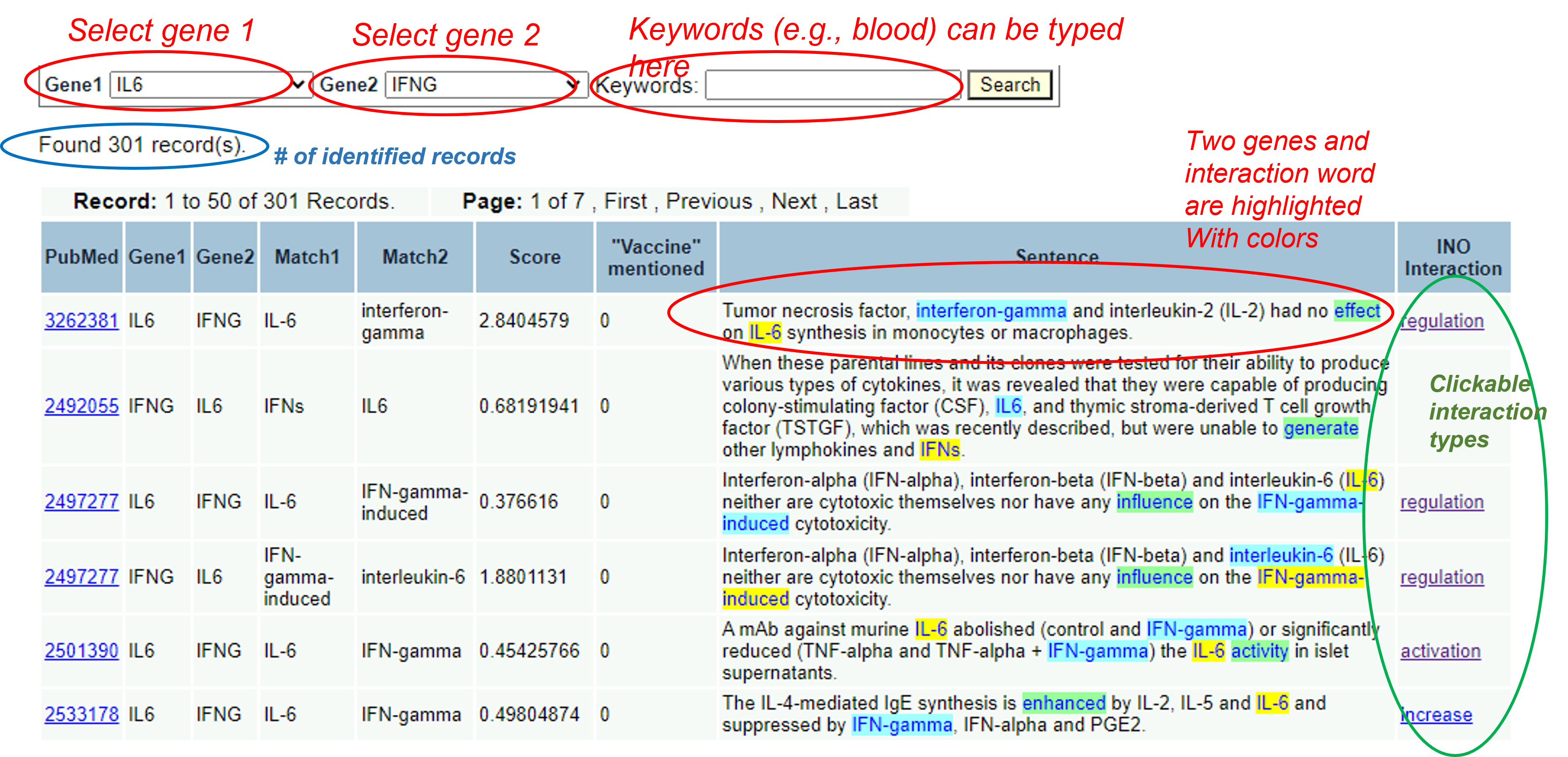
For more examples of GenePair analysis, you can click the following searching criterioa and find out the results:
- Interactions between IL6 and IFNG.
- Interactions between IL6 and IFNG with keyword "blood".
- Interactions between TLR4 and TNF.
- Interactions between TLR4 and IL10.
- Interactions between TLR4 and IFNG.
Note that the Interaction Network Ontology (INO) interaction keywords on the rightside of the above result page can be clicked to link to the interaction type definition and hierarchy shown on the Ontobee page, as demonstrated here:
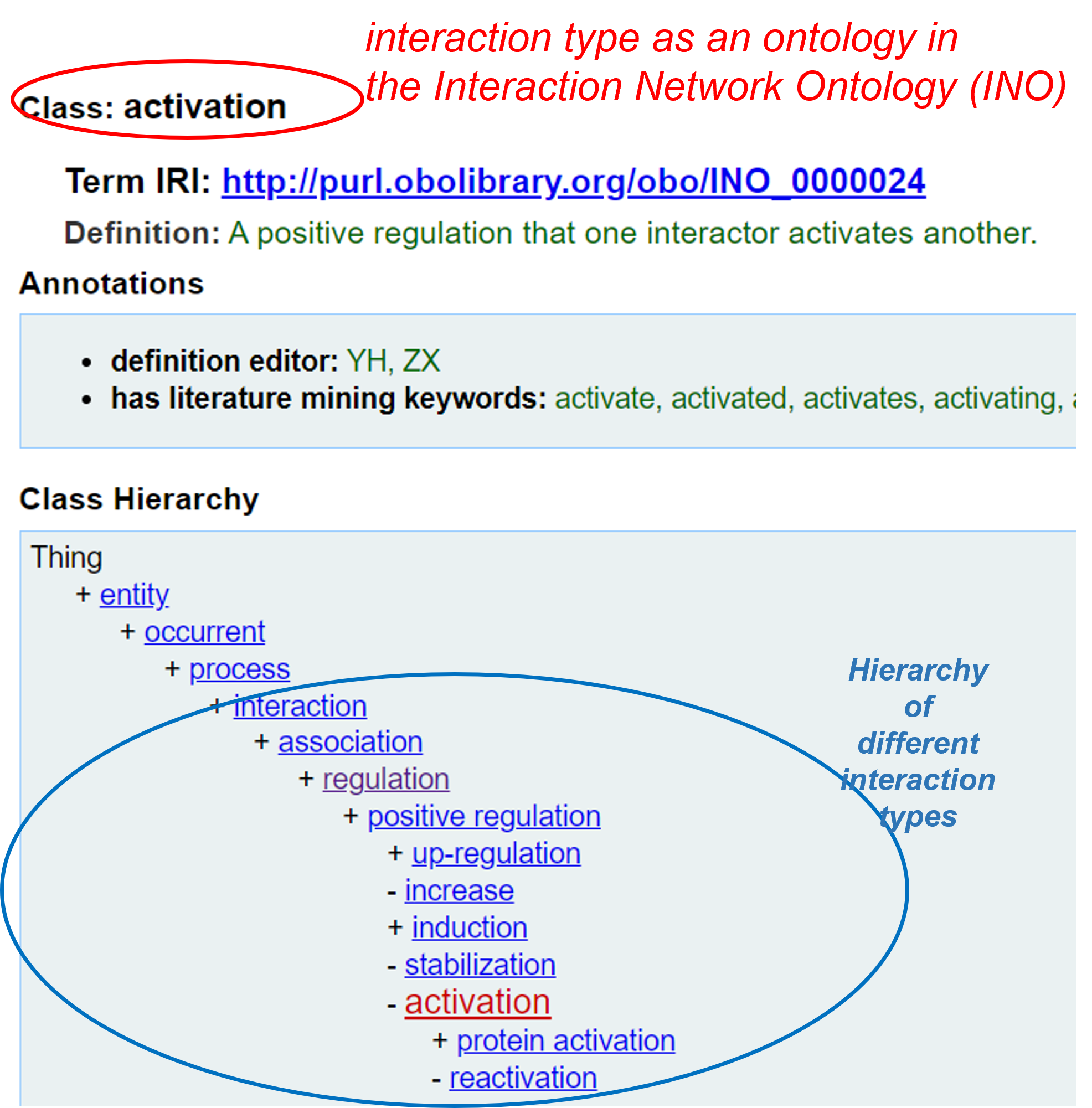

More information about how the INO ontology can help the literature mining and data analysis can be found in the following publications:
- Hur J, Özgür A, Xiang Z, and He Y. Development and application of an interaction network ontology for literature mining of vaccine-associated gene-gene interactions. Journal of Biomedical Semantics. 2015, 6:2. PMID: 25785184.
- Özgür A, Hur J, and He Y. The Interaction Network Ontology-supported modeling and mining of complex interactions represented with multiple keywords in biomedical literature. BioData Mining. 2016, 9:41. PMID: 28031747.
In addition to the INO, we have also applied other ontologies, including the Vaccine Ontology (VO), to study gene interaction network. The following papers demonstrate the usage of the VO for vaccine-related gene interaction network studies:
- Hur J, Xiang Z, Feldman EL, He Y. Ontology-based Brucella vaccine literature indexing and systematic analysis of gene-vaccine association network. BMC Immunology. 12(1):49 2011 Aug 26. PMID: 21871085. PMCID: PMC3180695.
- Ozgur A, Xiang Z, Radev D, and He Y. Mining of vaccine-associated IFN-γ gene interaction networks using the Vaccine Ontology. Journal of Biomedical Semantics. 2011, 2(Suppl 2):S8. PMID: 21624163.
- Hur J, Özgür A, Xiang Z, He Y. Identification of fever and vaccine-associated gene interaction networks using ontology-based literature mining. Journal of Biomedical Semantics. 2012, 3:18. PMID: 23256563.
- Hur J, Özgür A, Xiang Z, and He Y. Development and application of an interaction network ontology for literature mining of vaccine-associated gene-gene interactions. Journal of Biomedical Semantics. 2015, 6:2. PMID: 25785184.
Dynamic Dignet Query
The program is easy to run. Simply type a keyword(s), for example, "diabetes", "Brucella", or vaccine":
For example, below is a screenshot showing the searching for the typed keyword "vaccine":

Here are the results:
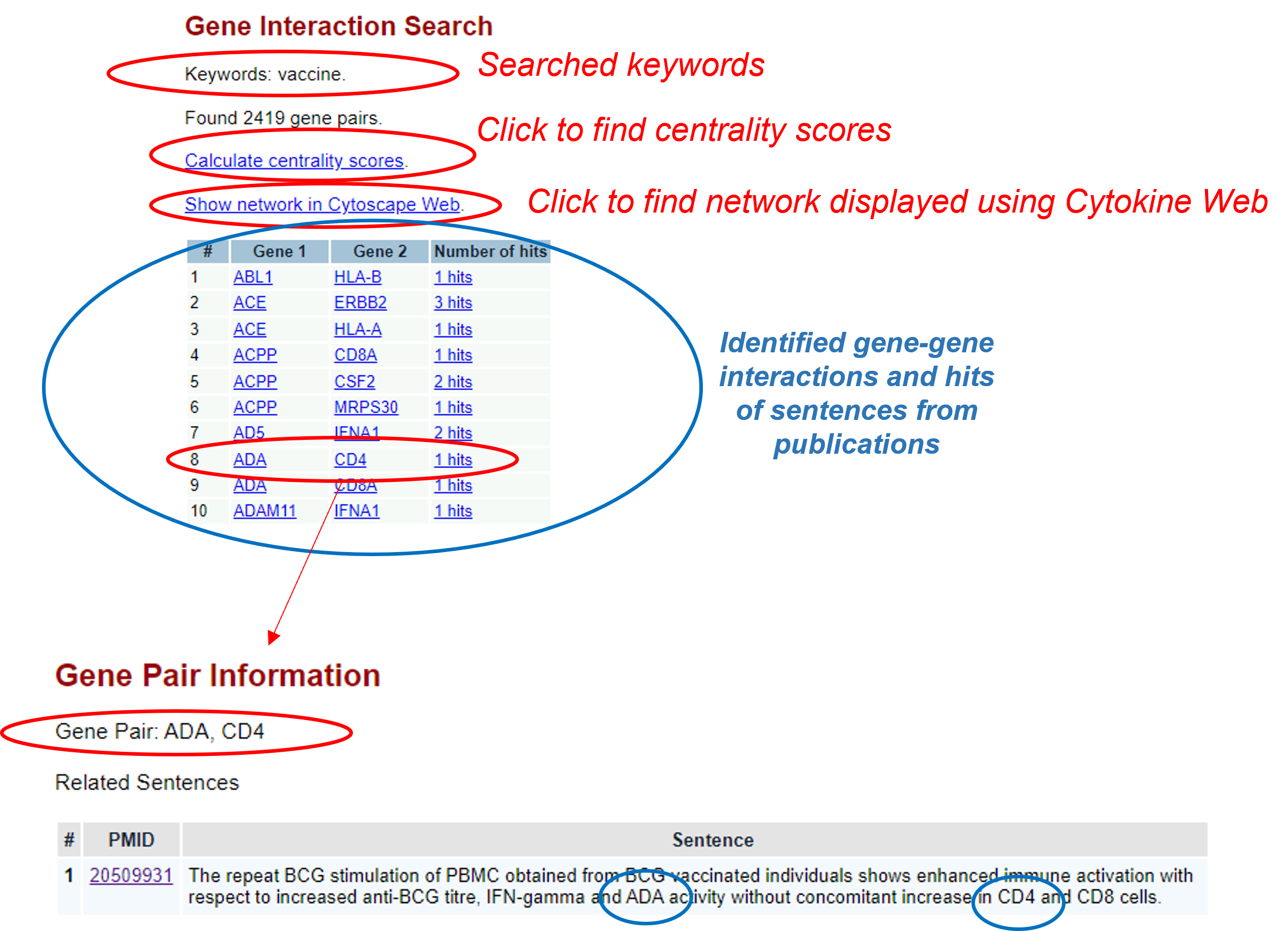
On the results pages, you can click different highlights and get different results as illustrated above.
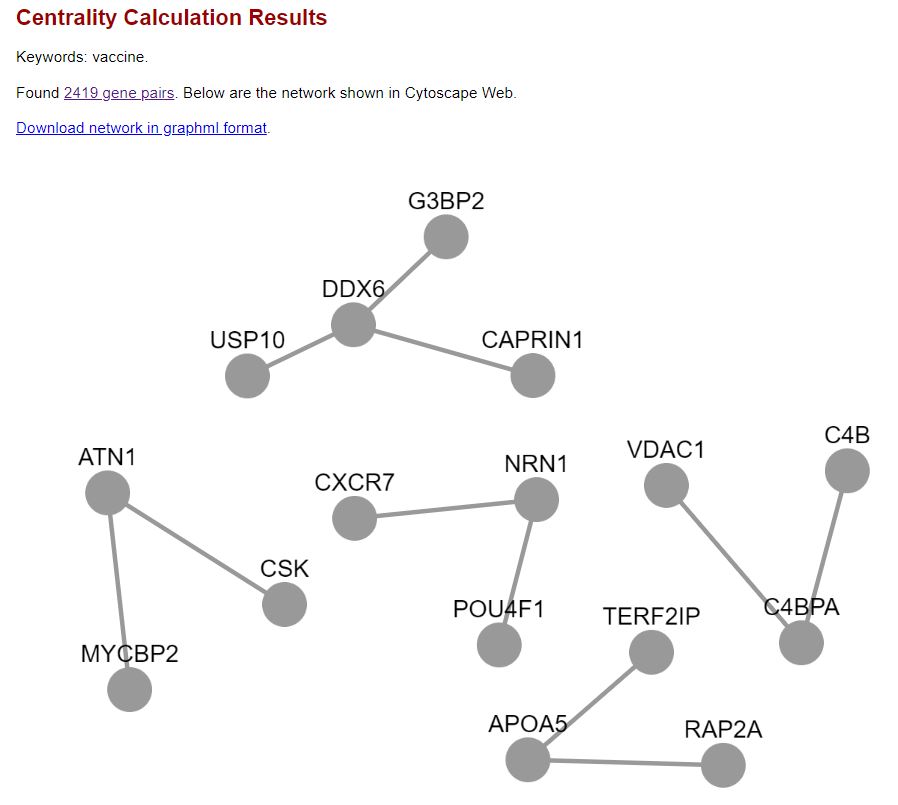
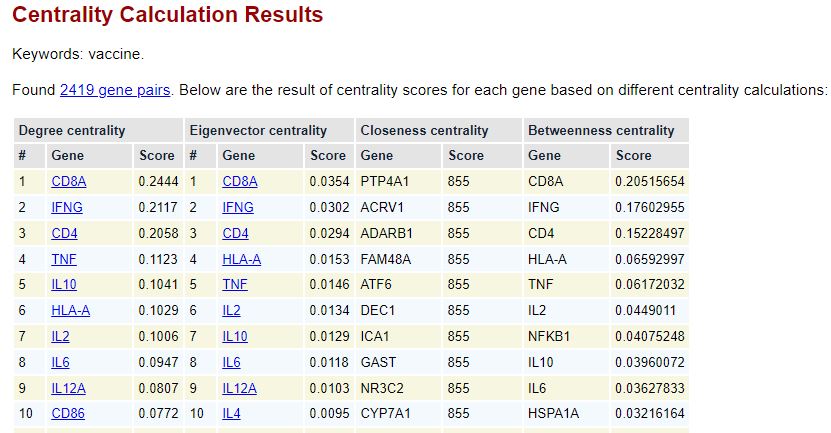
Below is the page screenshot showing the centrality calculation results:
Below is the page screenshot showing the high level graph network of the interconnected genes, which is generated using the Cytoscape Web technology:
Below is the page screenshot showing a specific portion of the graph network of the interconnected genes:
For more examples of Dignet gene interaction analysis, Choose and Click:
Note: The Ignet engine may have not yet processed most recent abstracts, so most recent results published in PubMed may not show up.
Please feel free to Contact Us for any suggestions, comments, and collaborations. Thank you!